A Step-by-Step Tutorial on sgRNA Design Using Benchling
#Benchling #CRISPR #Crisprcas9 #sgRNA
Welcome to our comprehensive guide on designing sgRNA with Benchling! I'm Elif Hangul, a dedicated high school student with a passion for gene editing. If you're curious about my current projects and what I'm up to, feel free to explore the links I've provided.
Now, let's dive into the exciting world of sgRNA design!
If missed a step and didn’t understand during the article, check out the video!! : https://www.loom.com/share/266119c7533047a9b69e4bb6ab7bc9cc?sid=696b66c8-fe20-4f51-93b6-2c4ddf792207
Gene Knock-out
A gene knockout is a genetic technique used to inactivate or "knock out" a specific gene in an organism. This process involves disrupting the normal function of a gene by introducing mutations that render it non-functional. Gene knockout remains a vital approach for understanding biology. By disrupting a specific gene, scientists can discover the effects and functions of that gene.
The CRISPR-Cas9 system, with the help of sgRNAs (single-guide RNAs), is a powerful and widely used method for creating gene knockouts. Here's a breakdown of how it works:
sgRNA Design:
sgRNAs are short RNA molecules designed to be complementary to a specific target gene sequence.
These sequences guide the Cas9 enzyme (a molecular scissors) to the precise location within the genome where the gene of interest is located.
Cas9 Enzyme Action:
Cas9 is an endonuclease enzyme that, when guided by the sgRNA, binds to the target DNA sequence.
Once bound, Cas9 induces a double-strand break in the DNA at the specified location.
DNA Repair Mechanisms:
When the DNA is cut, the cell's natural repair mechanisms come into play.
There are two primary repair pathways: Non-Homologous End Joining (NHEJ) and Homology-Directed Repair (HDR).
In NHEJ, the cell attempts to quickly repair the break, often introducing small insertions or deletions (indels) in the process. This can result in frameshift mutations that lead to a non-functional gene product. Non-homologous end-joining (NHEJ) is the best known and probably the most common repair mechanism that can make changes (mutations) in the DNA.
Functional Gene Knockout:
The introduced mutations disrupt the normal reading frame of the gene, preventing the synthesis of a functional protein.
As a result, the targeted gene is effectively "knocked out" or rendered non-functional.
Using sgRNAs in the CRISPR-Cas9 system provides researchers with a precise and efficient method for creating gene knockouts, enabling the study of gene functions, disease mechanisms, and potential therapeutic targets.
In this article, we will delve into the essential process of designing sgRNAs—a fundamental step in initiating CRISPR gene knockouts. This crucial step marks the inception of every experiment, serving as the cornerstone for precision gene editing and advancing scientific exploration.
Selecting the Target Gene
To commence, I recommend selecting a particular research paper detailing CRISPR-Cas9 treatment, preferably one related to a specific disease. This approach offers a valuable opportunity to gain a deeper understanding of the research and experimental processes. Within these research papers, the method section often provides insight into the target genes and sgRNAs used—allowing you to replicate and build upon their methodologies for your own investigations.
I opted for the article titled "CRISPR-Cas9 In Vivo Gene Editing for Transthyretin Amyloidosis." Amyloidosis is a condition characterized by the abnormal accumulation of amyloid proteins, leading to various organ dysfunctions. This disease holds significance due to its impact on vital organs and the potential for groundbreaking treatments through advanced gene-editing techniques. Moving forward, my next endeavor was to locate the pertinent genetic information.
As mentioned earlier, the methods section typically furnishes crucial information. In the case of the research paper I consulted, it specified the target gene for amyloidosis as the TTR gene. The provided sgRNA was designed to target the TTR sequence AAAGGCUGCUGAUGACACCU (in the human genome build hg38, located on chromosome 18: 31592987–31593007). Armed with this information, my next step involves understanding how to precisely design this sgRNA for the specified gene.
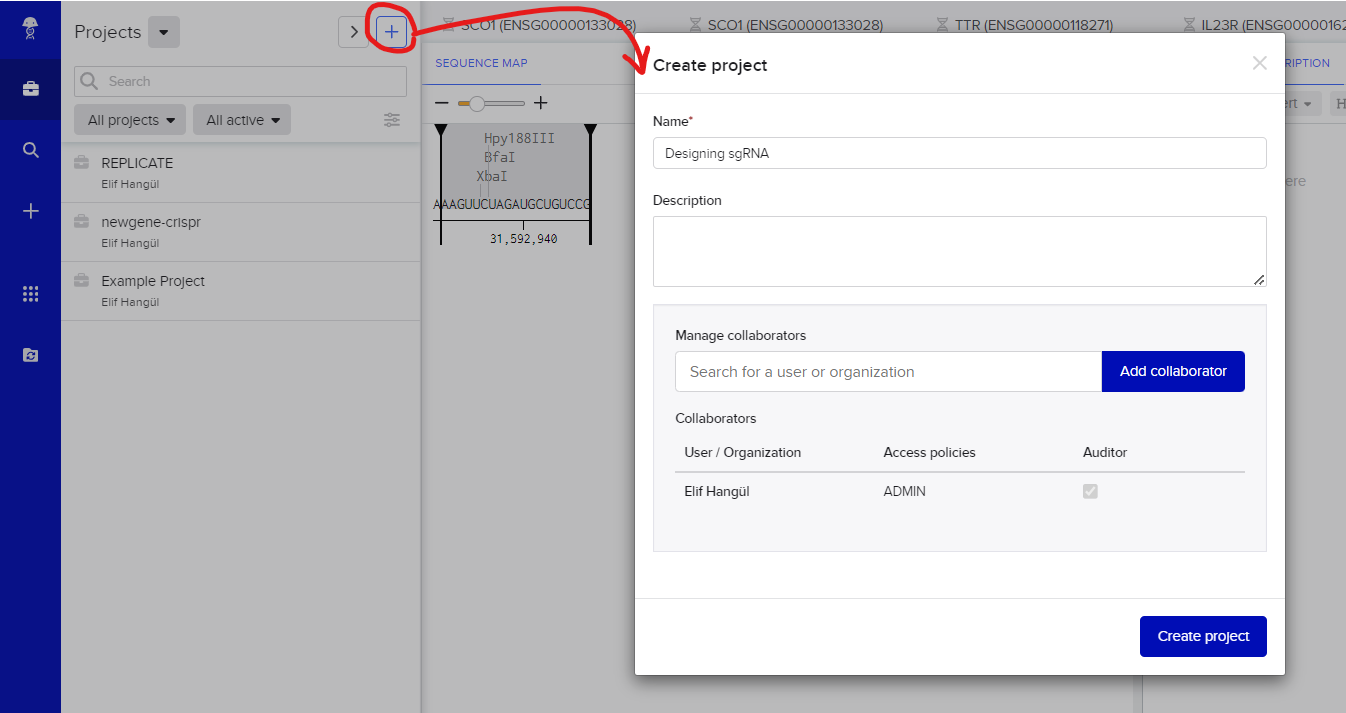
As our initial action on Benchling, we embark on creating a project. I've chosen to name it "Designing sgRNA," but feel free to customize the project title to align with your specific focus and objectives.
Next, navigate to your project screen, click the 'Add' button once more, and select 'CRISPR + CRISPR Guides' from the available options. This action will lead you to the screen displayed below.
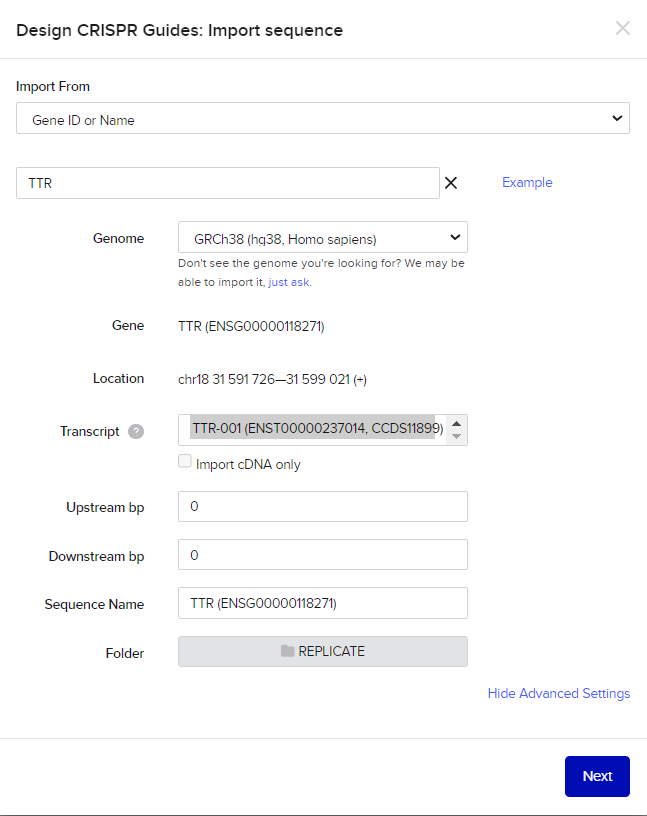
If you've selected your own research paper and gene, input the specific gene name. If you're following my example, enter "TTR" for the targeted gene. Below that, select "hg38" for the human gene. Additionally, click on "Show Advanced Settings" in the transcript section. Ensure you opt for displaying all transcripts to make an informed selection that aligns with your preferences.
For the guide parameters, leave the settings unchanged. Proceed by selecting "Finish" to conclude this step in the process.
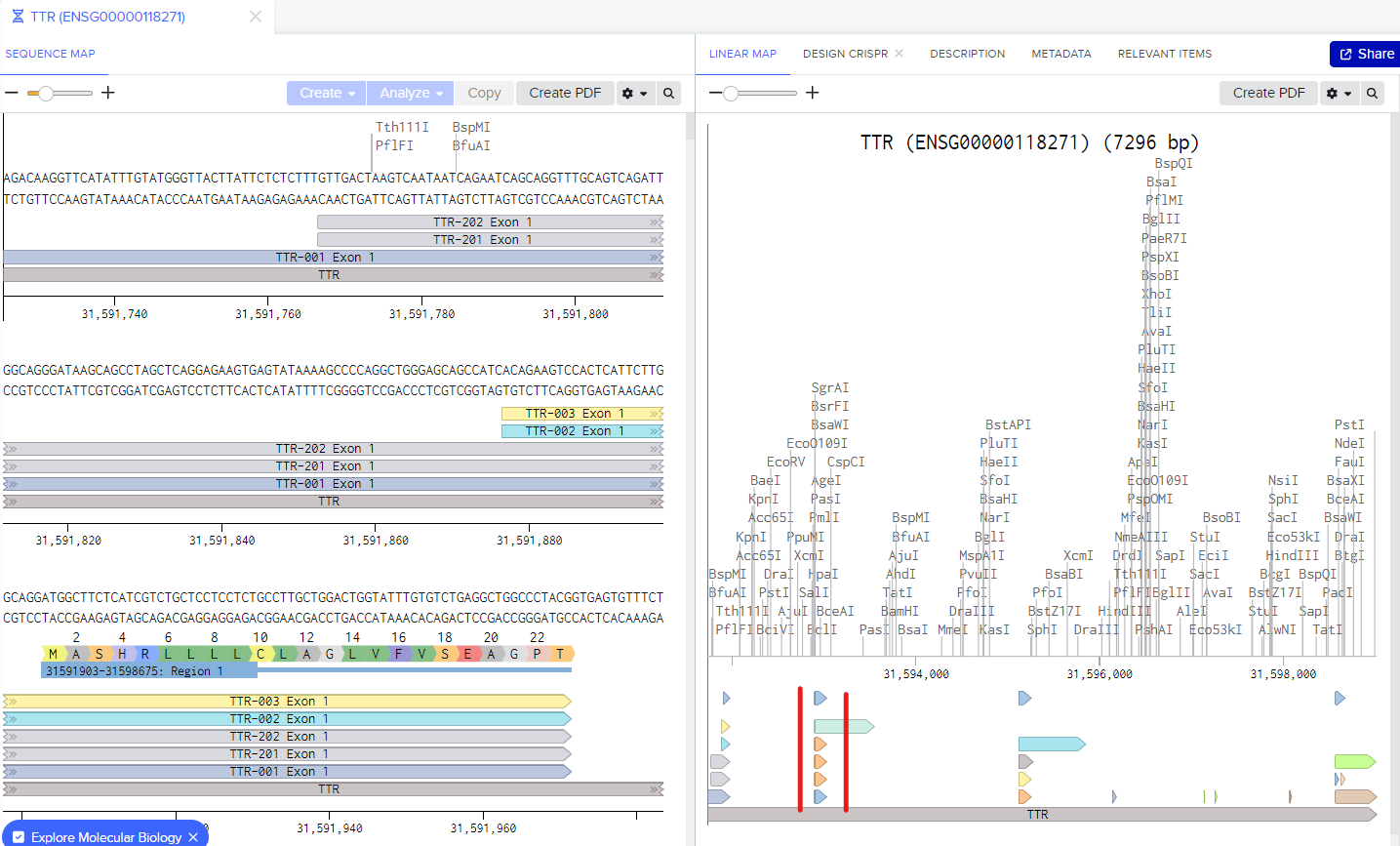
In the following step, you'll encounter the screen below. To proceed with the linear map, pinpoint a specific exon that corresponds to the desired gene section for your knockout. Each arrow block signifies a transcript of those exons. It's crucial to opt for the exon with the highest number of transcripts; selecting one with only one transcript might allow the cell to continue producing the proteins.
In my case, the optimal exon is exon 2 of the TTR gene. Click on the arrow block corresponding to your selection, and on the left, you'll observe the gene part you've chosen.
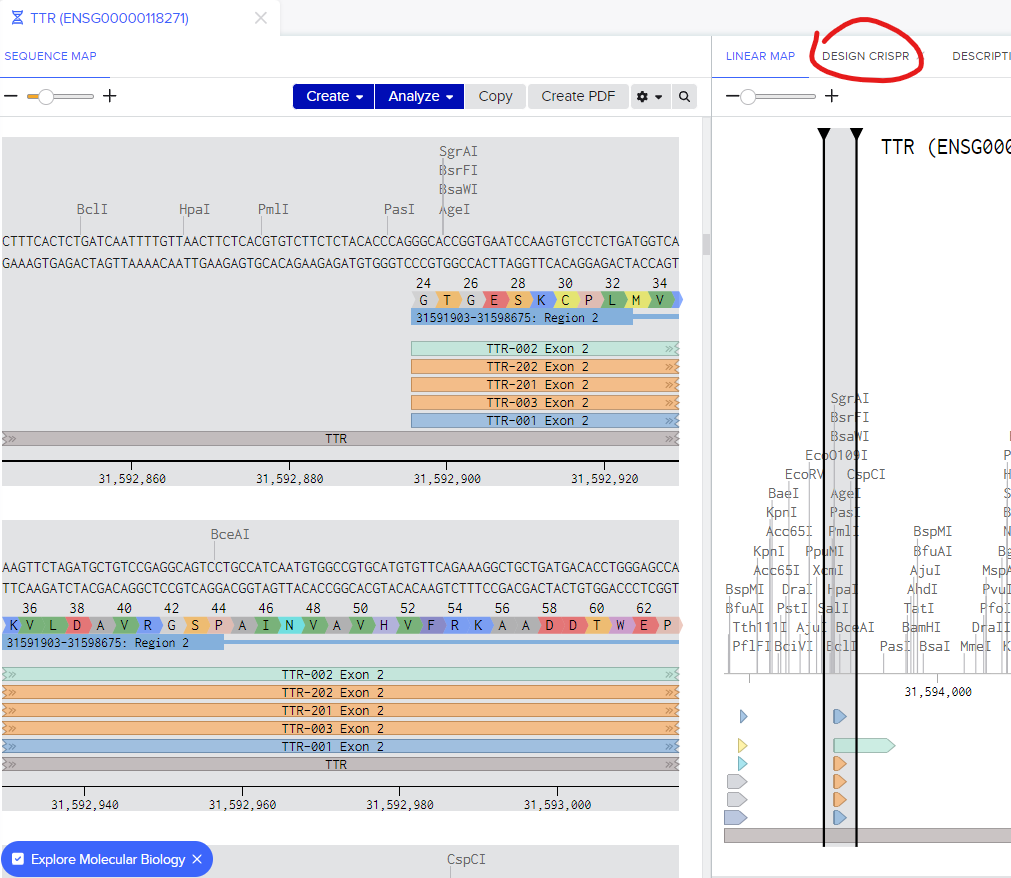
Having selected the appropriate exon with the highest number of transcripts for your gene, proceed to the "Design CRISPR" section. You'll notice that the targeted region corresponds to the exon you've chosen. Click on the "Add" button to explore the potential guide sequences available for your selection.
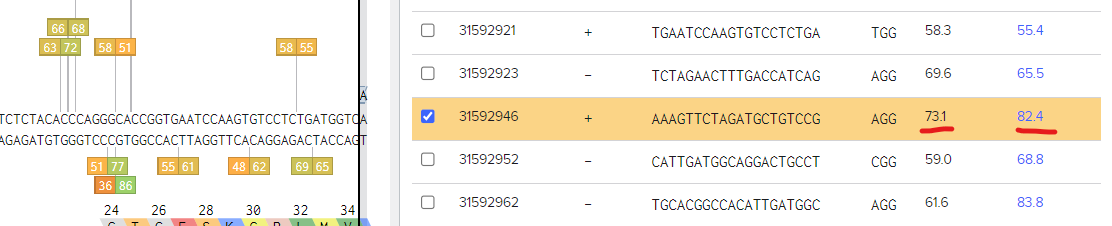
Now, we approach the critical phase of designing sgRNAs. Within the transcripts of the targeted region you've selected, you'll encounter a list of guide sequences. Each guide sequence is accompanied by both on-target and off-target scores, ranging from 0 to 100, where higher scores are preferable. Aim for values equal to or above 60. It's advisable to explore multiple options and select the ones that best align with the requirements of your experiment.
Fortunate news! We've stumbled upon an exceptional target guide sequence boasting impressive scores of 73.1 for on-target and 82.4 for off-target. This makes it the optimal choice for achieving a successful knockout in your experiment.
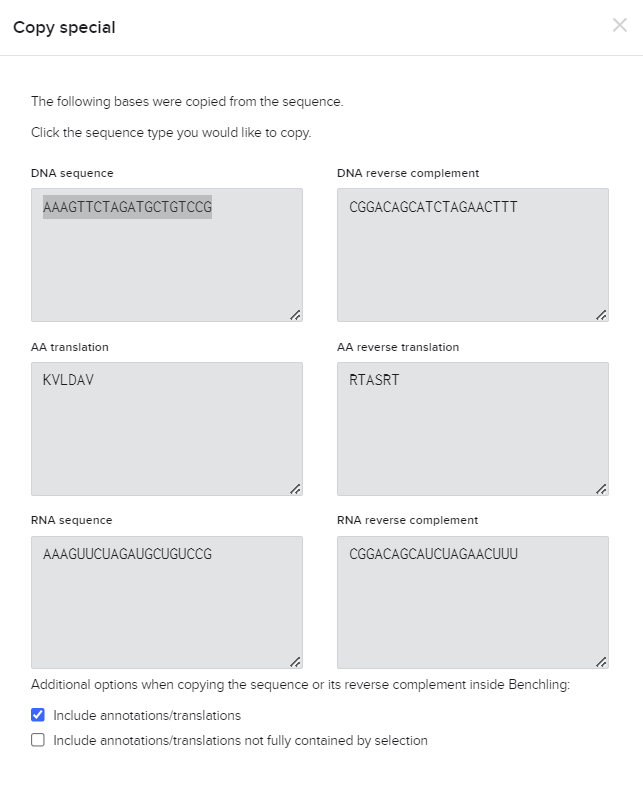
Now, simply click on the guide sequence on the left, specifying the targeted region. Choose "Copy," and you'll be directed to the following screen.
You've successfully copied the desired sgRNA sequence for your experiment! Now, for the next steps, consider ordering the sgRNA for your experiment. Reach out to a reliable supplier or facility equipped for custom sgRNA synthesis services. Provide them with the copied sequence, ensuring accurate specifications.
Keep in mind any additional requirements or modifications based on your experimental needs. Once obtained, you'll be ready to proceed with the gene knockout experiment using this meticulously designed sgRNA.
Thank you for exploring this guide! I trust you found it enjoyable and informative. Should you have any questions or seek further clarification, feel free to reach out to me via my linked LinkedIn profile mentioned at the beginning or through email at elifhangul06@gmail.com . Stay curious, stay tuned, and happy experimenting!