Exploring the Depths of Bioinformatics: Unraveling the Process Part 1
#geneediting #bioinformatics #rprogramminglanguage
Dear readers,
Welcome to this exploration of the fascinating world of bioinformatics! In this article, I aim to provide you a guide to learning bioinformatics, tailored specifically for beginners. As someone navigating the intricate landscape of molecular biology, my journey begins with the crucial step of selecting a reputable gene editing company. By delving into the tools and techniques employed by these companies, we can lay the foundation for a comprehensive understanding of bioinformatics methods. Join me on this educational adventure as we embark on the path to unravel the intricacies of gene editing and bioinformatics.
Why Bioinformatics
Bioinformatics, at its core, is a multidisciplinary field that combines biology, computer science, and statistics to extract meaningful insights from biological data. It involves the application of computational techniques and algorithms to analyze and interpret vast datasets derived from biological research.
In the realm of molecular biology and genetics, where the complexity of data has surged in recent years, bioinformatics plays a pivotal role. It facilitates the organization, analysis, and interpretation of biological information, ranging from DNA sequences to protein structures. This integration of computational tools into biological research has revolutionized our ability to uncover hidden patterns, make predictions, and derive valuable knowledge from the vast ocean of biological data.
The importance of bioinformatics lies in its capacity to accelerate research and enhance our understanding of complex biological processes. It aids in identifying genes associated with diseases, predicting protein structures and functions, and even designing novel drugs. Bioinformatics tools enable researchers to navigate through the intricacies of genomics, transcriptomics, and proteomics, making it an indispensable ally in advancing scientific discoveries and medical breakthroughs.
How Did I Start
Embarking on a journey into bioinformatics, one of the initial and essential steps is gaining insights into how leading companies leverage bioinformatics in their operations. This preliminary understanding serves as a foundation for several key reasons:
Real-world Application: Companies at the forefront of bioinformatics apply these tools and methodologies to solve practical challenges in the field of genetics, genomics, and molecular biology. By examining their practices, we can gain exposure to the real-world applications of bioinformatics.
Technological Landscape: Bioinformatics tools and technologies are continually evolving. By observing how companies incorporate the latest advancements, we can stay abreast of the dynamic technological landscape. This awareness is crucial for any one of us aspiring to contribute meaningfully to the field, as it helps in anticipating trends and staying adaptable to emerging technologies.
Networking and Collaboration Opportunities: Understanding how companies integrate bioinformatics opens doors to potential networking and collaboration opportunities. Familiarity with industry practices allows us to engage in more informed discussions with professionals in the field, enhancing our ability to collaborate effectively and contribute meaningfully to bioinformatics projects.
Selecting a suitable company may initially appear challenging due to the multitude of options available. In my case, I opted for Sangamo Therapeutics, drawn to their innovative applications like CAR-T cell therapy and the utilization of zinc proteins. It's essential to recognize that the ideal choice varies for each individual. By researching companies aligned with your interests and offering potential job opportunities, you can identify the one that best resonates with your career aspirations.
Sangamo Therapeutics
Sangamo Therapeutics, a prominent player in the biotechnology sector, is at the forefront of advancing genomic medicine and gene editing technologies. The company has garnered attention for its innovative approaches, particularly in the application of zinc finger proteins for precise and targeted gene editing. This novel technology involves engineering zinc finger nucleases (ZFNs) to specifically target desired DNA sequences, enabling meticulous modifications at the genetic level.
One notable area of Sangamo's work lies in the realm of CAR-T cell therapy, an emerging and promising field in the treatment of various diseases, notably certain forms of cancer. In the context of CAR-T cell therapy, Sangamo likely employs bioinformatics to analyze extensive genomic data. These analyses aid in optimizing the design of chimeric antigen receptor (CAR) T cells by identifying target genes, understanding gene expression patterns, and enhancing the therapeutic efficacy of these innovative therapies.
Beyond the laboratory, bioinformatics serves as an essential tool for Sangamo in the realm of data analysis and interpretation. Large-scale genomic data generated through various research initiatives require sophisticated computational approaches to extract meaningful insights. Sangamo likely leverages bioinformatics tools to navigate through this vast genomic landscape, identifying potential therapeutic targets, understanding the functional consequences of genetic modifications, and predicting the safety and efficacy of their interventions.
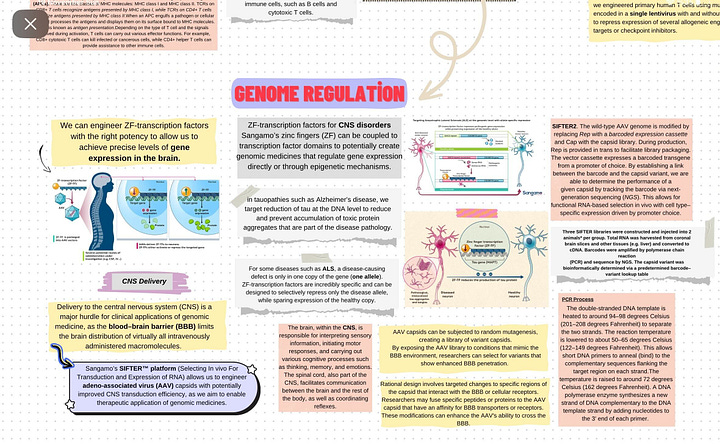
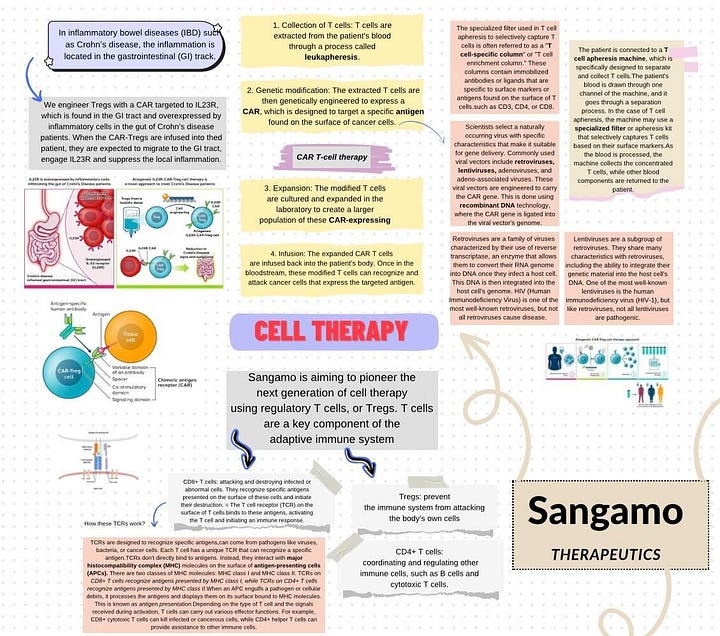
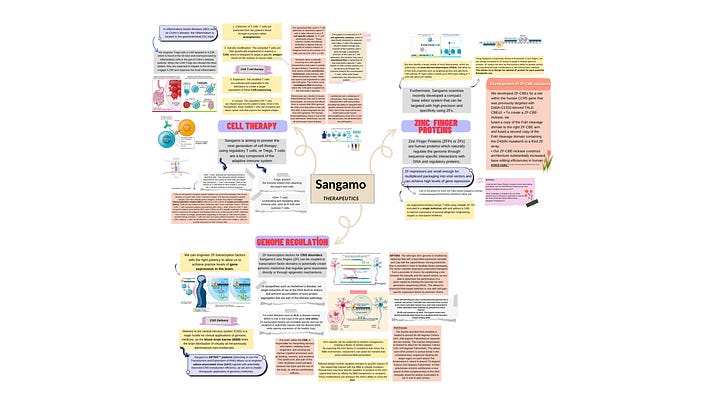




After dedicating a two-week period to thorough research on Sangamo and its applications (depends on your availability), I have synthesized my findings into a comprehensive mind map. To explore the intricacies further, you can click [here]. Utilizing a mind map proved to be an optimal approach, ensuring a clear and organized understanding of the subject matter. By categorizing each application separately, this visual representation facilitates a streamlined process, allowing you to select a specific area for more in-depth exploration in the further steps.
R Programming Language
In bioinformatics, we turn to R for its data prowess and statistical might. It's our go-to language for unravelling the intricacies of biological data, offering a rich toolbox of libraries designed just for us. With R, we seamlessly crunch large-scale genomic data, visualize complex datasets, and unearth insights from differential expression analyses. Its open-source spirit fuels collaboration, letting us share and refine tools worldwide. R isn't just a language; it's our key to unlocking the secrets within genomics and proteomics, a dynamic force in the ever-expanding realm of bioinformatics.
How To Learn It
Given the diversity in learning styles and abilities, my suggestion is to discover what works best for you. Explore various approaches to mastering the R language and tailor them to suit your preferences and objectives in bioinformatics. In this article, I'll share the method I employed in learning, offering insights that might resonate with your own learning journey.
Youtube Tutorials: I discovered that my most effective learning approach involves watching tutorials on YouTube first and then replicating the process independently. One standout channel, Bioinformagician, provides exceptional R language tutorials for bioinformatics. The creator, a young and passionate individual, makes the learning experience both engaging and straightforward. The tutorials meticulously cover every aspect, leaving no detail untouched. This method has proven remarkably easy for me to grasp complex concepts and execute them confidently. For those seeking a dynamic and thorough learning experience in R for bioinformatics, I highly recommend checking out Bioinformagician's tutorials [here].
5 down 19 to go
The initial phase of this bioinformatics learning guide encompasses the first five videos of the series. In subsequent articles, we will delve into the further steps, exploring advanced topics and expanding our knowledge in the fascinating realm of bioinformatics.
Let's kickstart our learning journey! Before diving into specific research cases, we'll begin by focusing on breast cancer through a series of tutorials. In these sessions, you'll gain hands-on experience in downloading patient data directly from the NCBI website. This acquired skill will serve as a foundation for our later steps, enabling you to confidently select and explore your chosen disease in more detail.
In the next phase of our learning journey, we'll delve into the art of data manipulation. Building on the foundation of breast cancer data obtained from the NCBI website, you'll acquire the skills to craft various datasets and tables. This process is crucial for extracting and segregating biological information, setting the stage for conducting extensive and impactful experiments.
Now, let's dive into my personal favorite part – data visualization! In the upcoming tutorials, you'll learn the art of professionally visualizing biological data. These skills will empower you to create compelling graphics that can enhance the visual impact of your research papers and experimental reports. From intricate charts to insightful graphs, we'll explore a range of visualization techniques. Here's a sneak peek at some of the graphics we'll be crafting in the tutorials.
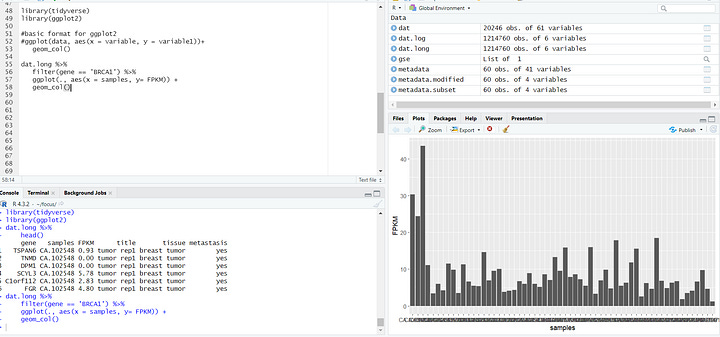
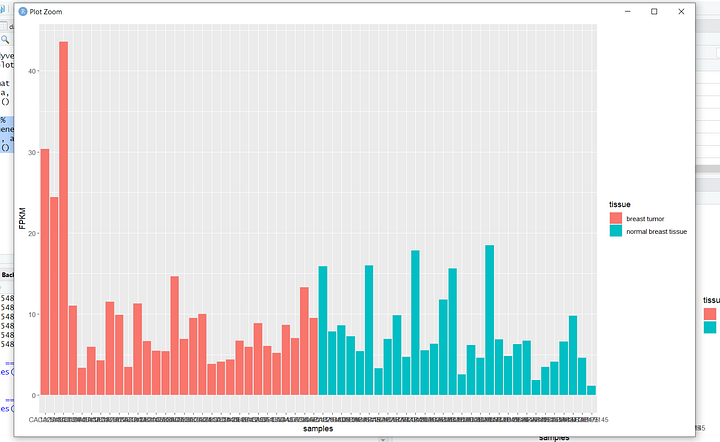
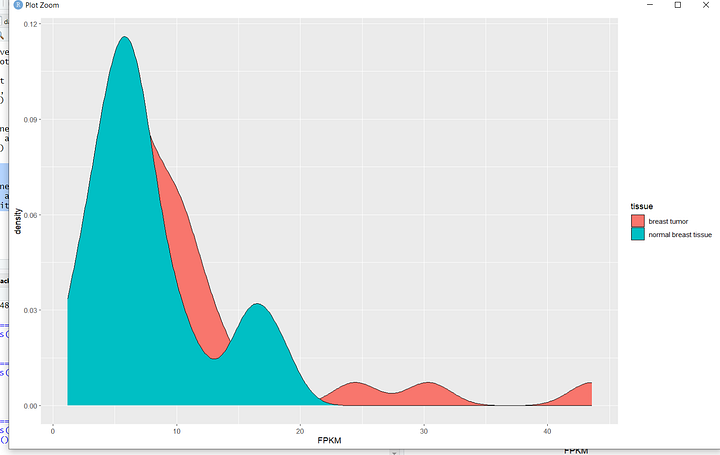
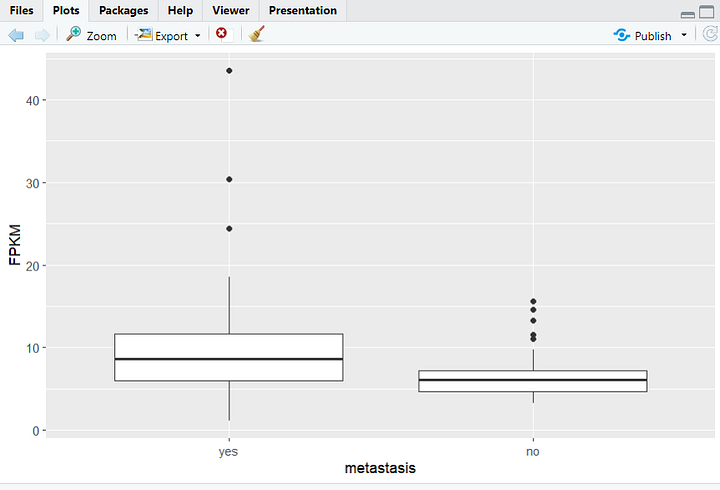
In the final phase of this learning series, we'll broaden our horizons by delving into a different type of data—SRA, accessible through the NCBI website. SRA stands for Sequence Read Archive, a repository that stores raw, high-throughput sequencing data. It includes vast amounts of genetic information from various organisms, providing a goldmine for researchers.
In the final installment of this tutorial series, we'll explore the differences between RNA-Seq normalization methods, with a specific focus on techniques like FPKM and RPKM. Understanding these normalization methods is paramount in the realm of bioinformatics, as they play a crucial role in ensuring accuracy and comparability across RNA-Seq datasets.
Before we venture into the intricacies of DESeq2 in the upcoming videos, it's crucial to take a moment to understand our learning. In the following sessions, we'll revisit and reinforce the concepts we've covered, ensuring a solid understanding of the fundamentals. This reflective pause offers the perfect opportunity to adapt our newfound knowledge into practical applications for your individual projects.
How to Personalize Our Learning Experience
Recall our initial research on a company? We immersed ourselves in exploring their applications, creating a structured mind map to comprehend their mechanisms. Now armed with insights into how they initiate experiments, let's use this knowledge to replicate and apply similar methodologies in our own projects.
My enthusiasm lies in CAR-T cell therapy, and upon reviewing my mind map, you'll notice a specific focus on Sangamo's application of this therapy for patients with Crohn's Disease. Recognizing the potential, I've decided to make this the cornerstone of my replicate project.
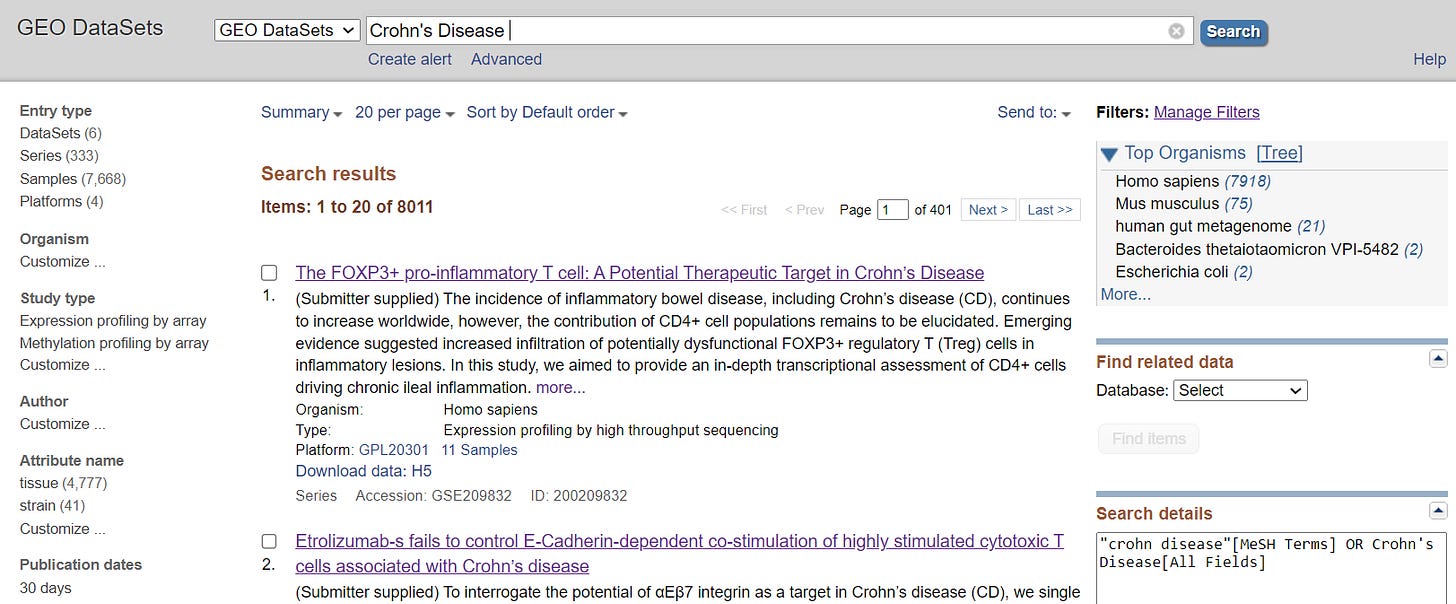
Once you have selected the disease you want to focus on, utilize the NCBI website to identify the appropriate dataset. Specifically, navigate to GEODatasets and enter the name of the chosen disease for relevant information.
You will encounter numerous datasets. One significant mistake I made was neglecting to verify the file type. If it's in H5 or another format, it becomes challenging to convert and read in R. The most preferable format is CSV, and you will learn how to handle CSV files in tutorials. I've spent considerable time attempting to convert tar (H5) files, and I must admit, it was quite traumatizing! Make sure to find the dataset that best fits your needs, and don't forget to confirm if it's in CSV format.
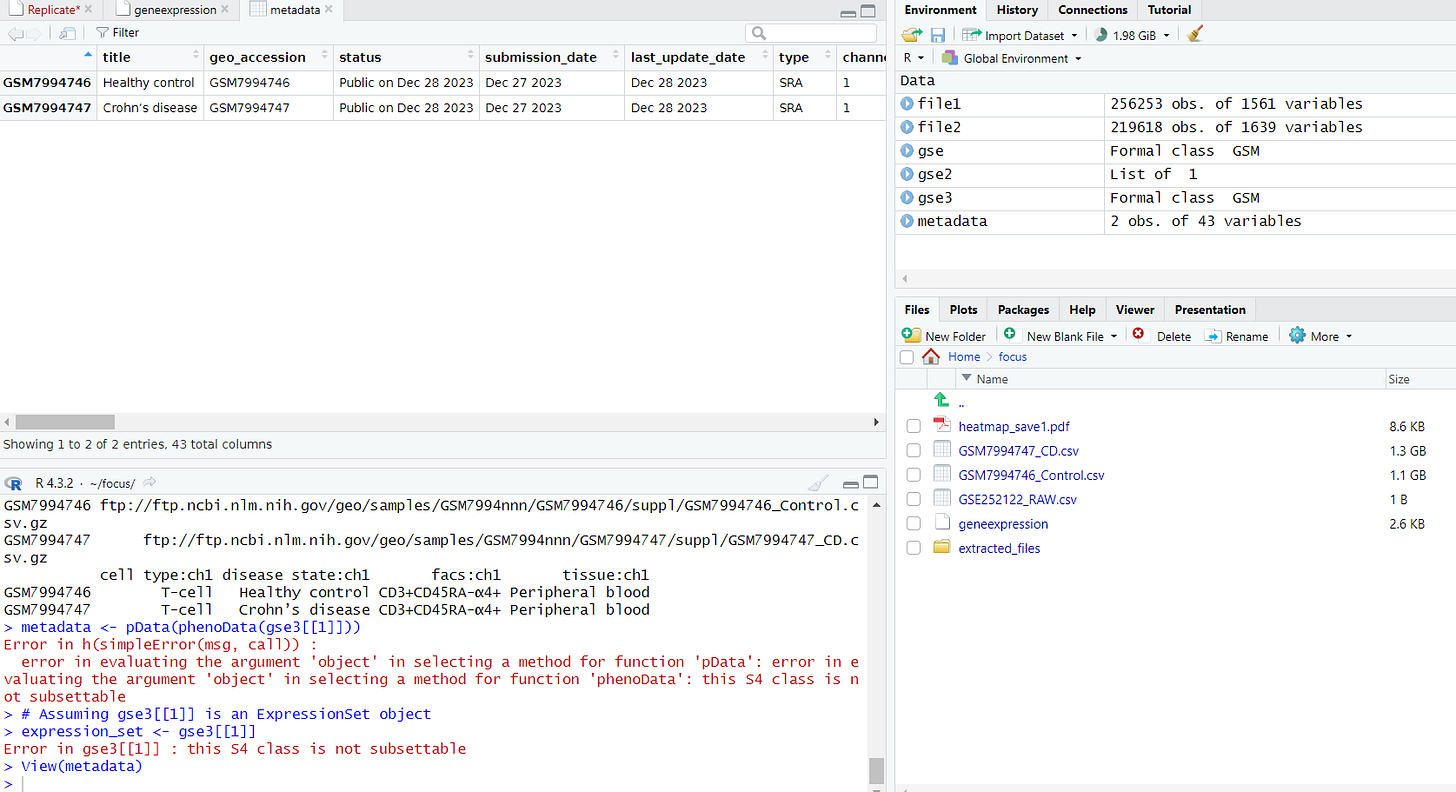
Following that, all you need to do is follow the steps you learned from the tutorials applicable to your dataset! While it may initially be challenging to adapt to the specifics of your dataset, with time, you'll gain a full understanding of what each step entails. Feel free to ask questions, including reaching out to me through the contact information provided at the end.
In my case, my dataset lacked numerous pieces of information necessary for creating visualizations. However, I addressed this by creating metadata that includes crucial details required for research papers. These tables have significantly streamlined the process of writing research papers and preparing reports. In your dataset, specifically check if there's an FPKM section in your metadata. If it exists, experiment with every visualization method you learn. This will undoubtedly enhance your experience and understanding of the data.
And that brings us to the end, dear readers! I hope you find this guide helpful. In upcoming articles, we'll delve deeper into bioinformatics and its applications. Stay tuned!
As mentioned earlier, feel free to reach out to me with any questions. Here is my contact information:
My Email: elifhangul06@gmail.com
My LinkedIn account: Click here!